Emran Ali, Owen Hudson, Justin Hand, and Sumyya Waliullah
Georgia ranks among the top three states in the nation in vegetable production. One of the most serious diseases in vegetable production in Georgia is Phytophthora blight, caused by the oomycete pathogen Phytophthora capsici. It is a water mold that attacks the roots, foliage, and fruit, causing root rot, crown rot, leaf lesions, fruit rot, and plant wilt (Fig.1). The disease affects peppers, squash, watermelon, cucumber, cantaloupe, and other vegetable crops.
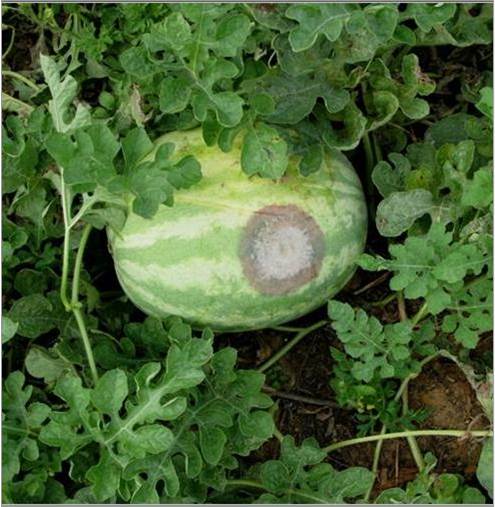

Fig. 1 Example of watermelon fruit rot caused by Phytophthora capsici (Photo credit: Dr. Pingsheng Ji)
The continuous rainfall in Georgia makes Phytophthora blight a widespread problem on vegetables. Because this pathogen produces spores (sporangia and zoospores) on the surface of diseased plant tissues, the spores can be easily washed out by splashing rain and can contaminate nearby irrigation sources like irrigation ponds or lakes.
Previous studies indicated that this pathogen can survive in irrigation water that may serve as an inoculum source. Due to a lack of efficient diagnosis systems, the production of vegetables is severely impacted by contaminated irrigation water.
Detection of P. capsici in irrigation water is difficult using traditional culture-based methods because of other microorganisms present in the environment, such as Pythium spp., which usually overgrow on culture media making P. capsici undetectable. To detect the presence of P. capsici spores in water sources (irrigation ponds, runoff, etc.), we developed a hand pump-based filter paper (8-10 µm) method that captured zoospores and was used to amplify DNA of the pathogen through a novel loop-mediated isothermal amplification (LAMP) assay designed for specific amplification of P. capsici (Fig. 2).
This method amplified and detected DNA from a concentration as low as 1.2 x zoospores/ml, which was 40 times more sensitive than conventional PCR. No cross-amplification was obtained when closely related species were tested.
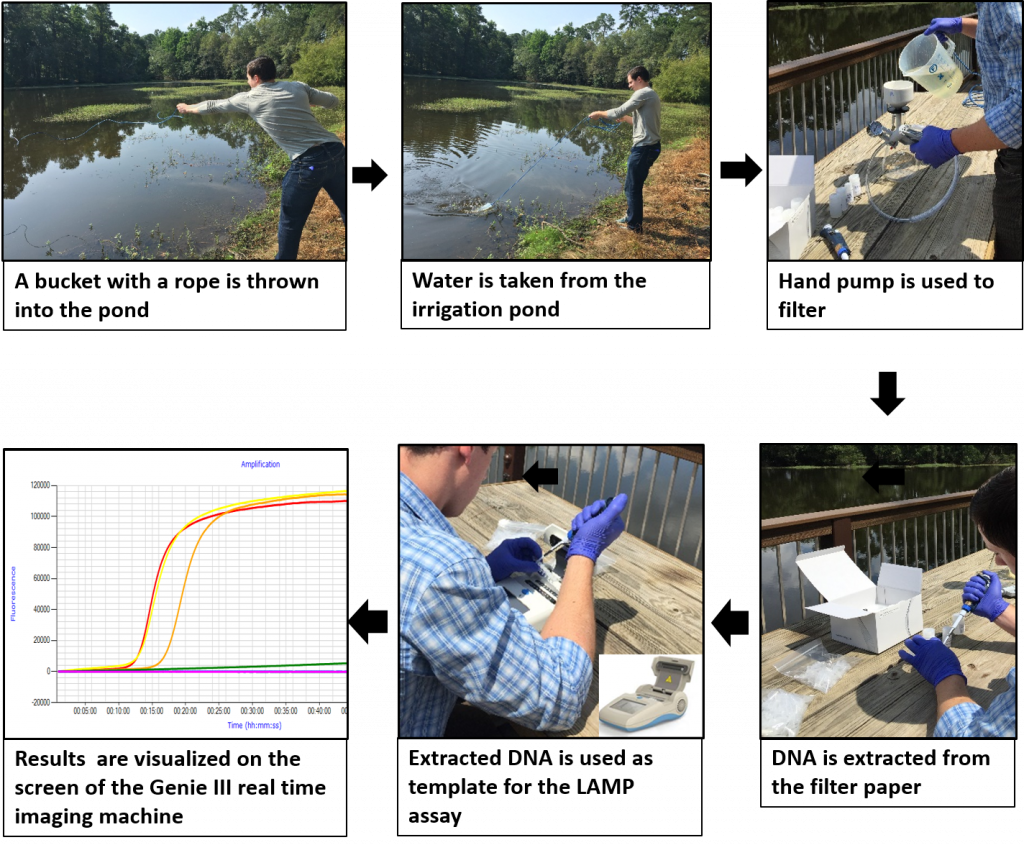
Fig. 2: Pictures showing the sampling and processing of recycled water for the detection of Phytophthora capsici in the field.
To validate our detection protocol, water samples from the field where P. capsici was suspected to be present was taken to test the designed method with a practical scenario. Out of the seven farms tested, three were positive for the presence of P. capsici using our hand pump filter paper-based LAMP assay. Only one farm was positive when using the conventional PCR assay (Table 1), showing LAMP to be a more sensitive assay for this method of testing irrigation water.
Table 1. Detection of irrigation water from Southern GA
| Pond name | County, State | Target crops | Filter paper-based LAMP detection | PCR Detection | History of Disease (Y/N) |
| P1 | Tift, GA | Vegetables | + | - | N |
| P2 | Tift, GA | Vegetables | – | – | N |
| P3 | Tift, GA | Vegetables | – | – | N |
| P4 | Tift, GA | Vegetables | + | + | N |
| P5 | Tift, GA | Vegetables | – | – | N |
| P6 | Tift, GA | Vegetables | + | – | N |
| P7 | Tift, GA | Vegetables | – | – | N |
This improved detection method will enable researchers and extension agents to directly utilize the protocol described here to detect P. capsici. spores from a water source in less than two hours. We hope that this will lead to an increase in awareness of using pond water as an irrigation source which will eventually improve disease management of P. capsici, reduce production cost and increase crop yield. This protocol could be adapted to other pathogens that reside, accumulate, or are dispersed in contaminated irrigation systems.
Moving forward, growers should have their irrigation sources like ponds tested for the presence of P. capsici. The Plant Molecular Diagnostic Laboratory, a lab service of the University of Georgia Department of Plant Pathology, is now providing P. capsici testing support for vegetable growers in Georgia. The clinic can accept water samples (generally 2 L water samples per site) to test for the presence of P. capsici. The tests currently available, their pricing, a submission form, and submission information are available at the MDL web page at https://site.caes.uga.edu/alimdl/
https://site.caes.uga.edu/alimdl/files/2018/08/Submission-form-MDL-latest-7-5-18.pdf
Samples can be shipped to the following address.
Plant Molecular Diagnostic Lab
Department of Plant Pathology
Tifton, CAES Campus
Plant Science Building
115 Coastal Way
Tifton, GA 31794
The contact information for questions, etc. from Dr. Ali are as follows:
229-386-7230
229-386-7285
Again, we would highly encourage you to take advantage of this service. If you have questions or need help, please contact your local county extension agent for additional information. It would be good to communicate with the lab so that they can expect the samples on the day of arrival.









